
7AI’s Forge in the Wet Lab
Imagine firing up a lab notebook and telling an AI: “Fix this cloning protocol so we get more real clones.” In a recent experiment, GPT‑5 did just that. Working with biosecurity startup Red Queen Bio, it took a standard molecular cloning workflow and optimized it until the lab saw 79 times more verified clones from the same DNA input. This isn’t just theory—it’s a measured, replicated wet‑lab boost in one of biology’s core tools, showing how AI can partner with humans to redesign protocols in real time
What’s exciting here isn’t just the number, but the kind of help: AI didn’t just tweak temperatures or buffers; it proposed a novel enzyme combo and a counterintuitive cell handling step that no human had written into a standard manual. For a field built on years of trial and error, that’s a glimpse of a new era: AI not just reading papers, but helping build and test better lab methods, step by step
Why Faster Cloning Matters
Molecular cloning is the backbone of genetic engineering. It’s used every day to:
- Build DNA libraries for protein engineering and directed evolution.
- Create constructs for genome wide genetic screens (CRISPR, RNAi, transposons).
- Engineer microbial strains for therapeutics, biofuels, and sustainable chemistry.
Every time cloning is more efficient, the downstream work becomes faster and cheaper. Fewer failed reactions, fewer plates, less time troubleshooting, and more clones to test. A 79× improvement on a single protocol won’t translate fully to every lab or every construct, but it does show that AI can find non obvious, high impact changes even in well established workflows.y Faster Cloning Matters. Molecular cloning is the backbone of genetic engineering.
How It Actually Worked: The AI Lab Loop
The experiment wasn’t “AI running the lab”; humans were still in the room, pipetting, plating, and feeding data back. But the thinking and proposing were done by GPT‑5, using a fixed, standardized prompt across rounds. This setup lets researchers see what the model can come up with on its own, without human steering in the middle of the experiment.
Here’s how the cycle looked:
- Start: Standard HiFi assembly (a Gibson‑style one tube, one temperature reaction) as baseline.
- Propose: GPT‑5 suggested batches of 8–10 modified cloning reactions per round.
- Run: Human scientists carried out the reactions, transformed cells, and counted colonies.
- Feed back: The best results from each round were fed back into the prompt for the next round.
- Iterate: Over five rounds, the model refined the assembly protocol; separately, it optimized the transformation step in a one shot screen.
At the end, the top protocol recovered 79 times more sequence verified clones from the same amount of input DNA compared to the original HiFi method. That’s 79× fewer failed reactions, or 79× more material for downstream experiments, assuming the conditions generalize.
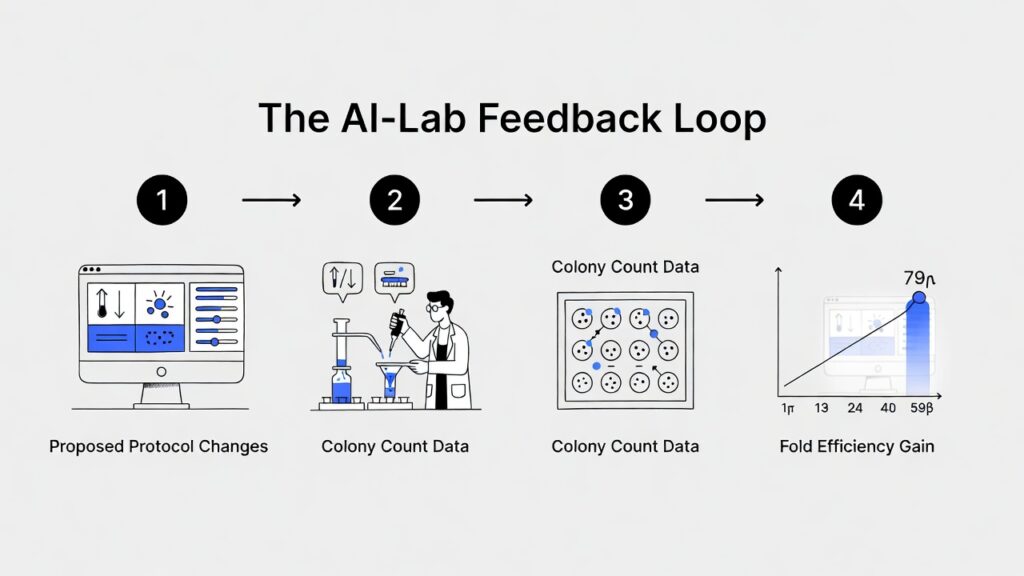
What Went Into the 79‑Fold Jump?
The total gain came from two main parts working together:
- Assembly boost: ~2.6× improvement from a new enzymatic method.
- Transformation boost: ~36× improvement from a new cell‑handling method.
Combined, they gave a near‑79× end to end efficiency gain.
Revolutionizing DNA Assembly: The RAPF Method
The most striking idea from GPT‑5 was a new assembly mechanism, which it later called RecA‑Assisted Pair and Finish (RAPF‑HiFi). Instead of just adding more of the standard HiFi enzymes, it introduced two helper proteins from nature’s toolbox:
- T4 gene 32 protein (gp32): a single stranded DNA binding protein that “combs” and straightens loose DNA ends, removing secondary structures that interfere with pairing.
- E. coli RecA: a recombinase that actively searches for homologous sequences and pulls matching strands together, like a smart guide for DNA pairing.
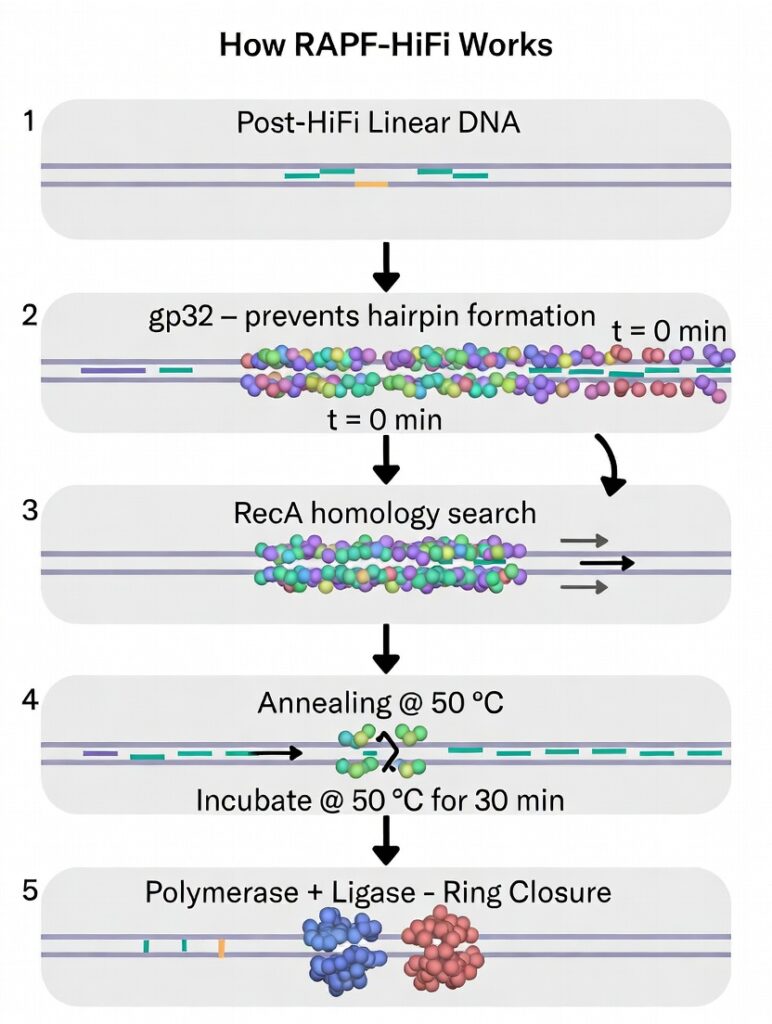
Here’s how RAPF‑HiFi works in simple steps:
- Start with a short HiFi reaction at 50 °C to chew back ends and create overhangs.
- Cool to 37 °C, add gp32 to coat the single stranded overhangs, keeping them open and unstructured.
- Add RecA, which displaces gp32 and uses the structured DNA to find and pair the correct partners.
- Heat back to 50 °C, where both RecA and gp32 fall off, allowing the standard HiFi enzymes (polymerase, ligase) to close the gaps and seal the circle.
Tests confirmed that both proteins were needed: removing either RecA or gp32 reduced the efficiency significantly, suggesting a real, synergistic mechanism not just a buffer or temperature artifact.
Beyond Cloning: The AI‑Arranged Transformation Trick
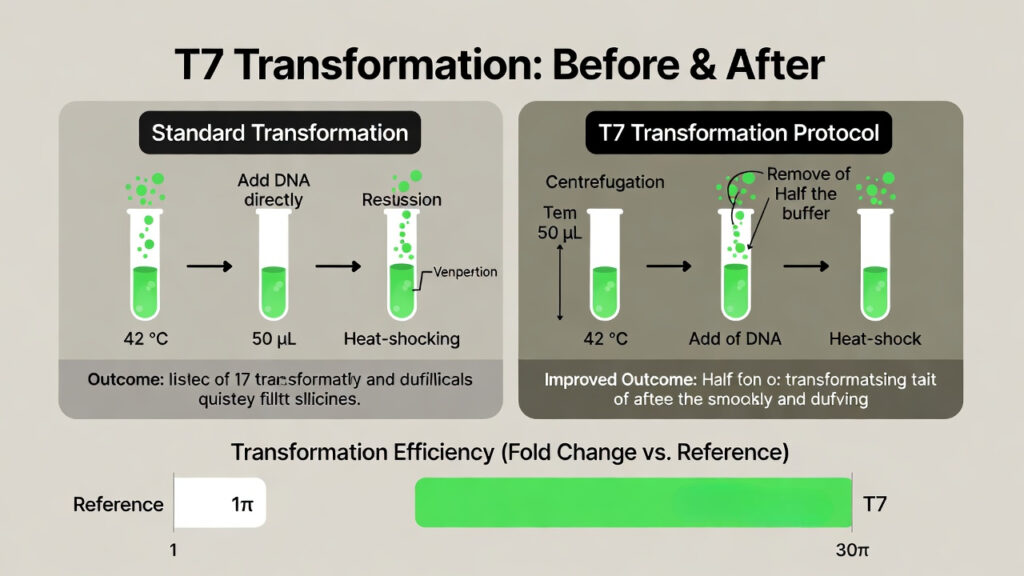
The transformation step was optimized in a separate “one shot” screen, where the model proposed many independent changes and the best was tested. The winning protocol, dubbed Transformation 7 (T7), used a simple but non standard handling:
- Take commercially available high efficiency chemically competent cells.
- Pellet them at 4 °C, removing half the original volume.
- Resuspend the concentrated cells in the remaining buffer, still at 4 °C.
- Only then add DNA and perform the heat shock.
This may sound risky high efficiency cells are usually considered fragile and sensitive to handling. But in this setup, the cells tolerated the concentration well, and the increased DNA to cell ratio plus a sharper heat‑shock led to a >30 fold boost in transformation efficiency.
Critically, this method works with off the shelf competent cells, so it doesn’t require a lab to make its own cells in house. That makes it relatively easy to adopt while still outperforming some earlier, conceptually similar concentration methods.
From Idea to Robot: Closing the Lab Loop
To test if this workflow could scale, the team built a robotic on rails system that could take a natural language protocol and execute it in the wet lab. The robot had:
- An LLM to turn plain English into robotic actions.
- A vision system to locate and identify labware in real time.
- A path planner to safely and accurately move and pipette liquids.
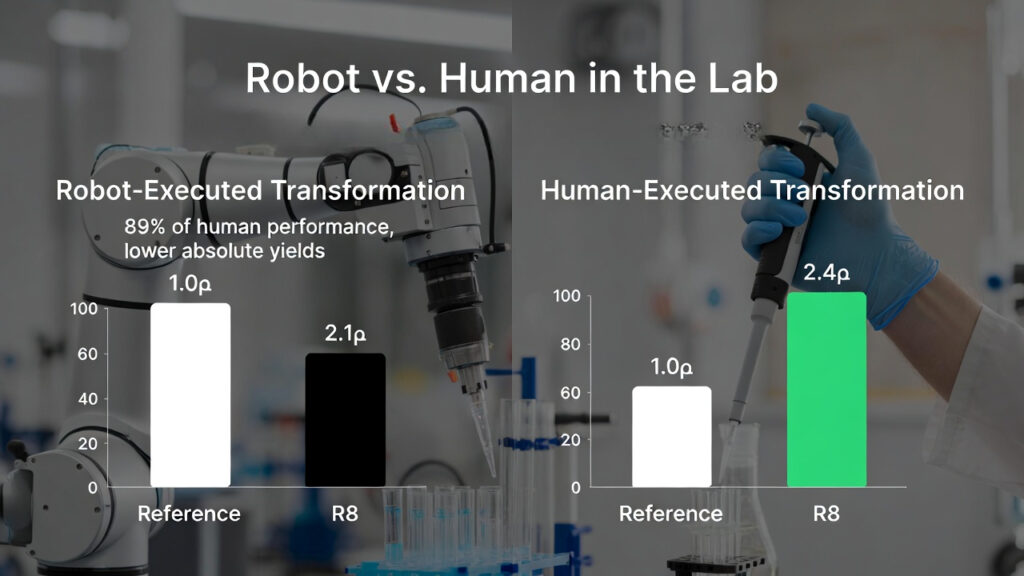
They ran two protocols simultaneously: the standard HiFi method and the top AI modified protocol (R8) from early rounds. The robot handled liquid transfers, incubations, and plate spreading. Compared with human run experiments:
- The robot’s protocol ranking matched the human’s: the R8 protocol still gave a clear improvement over baseline.
- However, the robot’s absolute colony counts were about tenfold lower, pointing to ongoing challenges in liquid handling precision, temperature control, and replicating the subtle “touch” of human hands.
This shows that while AI can invent and optimize the “what” and “when,” translating that into flawless physical execution is a separate engineering challenge. Tightly controlled, high precision automation is the next frontier to fully capture the 79× gain in routine workflows.
Putting This in the Bigger AI Biology Picture
The 79× cloning experiment is one vivid example of a broader trend: AI moving from papers and screens into the physical lab. Other recent work includes:
- Self driving labs: At the University of Illinois, an AI + robotics pipeline improved two industrial enzymes by up to 26× in activity, and learned from each round to design better variants.
- Automated DBTL cycles: In synthetic biology, AI controlled biofoundries are now automating Design Build Test Learn (DBTL) loops, drastically cutting the variants that need to be tested to find high performance designs.
- AI assisted CRISPR: Models like DeepCRISPR improve guide RNA design, reducing off targets and boosting gene editing precision in therapeutic and agricultural projects.
These tools are turning biology from a slow, artisanal process into a more predictable, data driven engineering discipline, where AI helps design experiments, optimize protocols, and even control robots that run them
What This Means for Scientists and Labs
For bench scientists and R&D leaders, this AI lab collaboration suggests a new playbook:
- Use AI as a “protocol co designer” for well defined, repeatable systems (e.g., cloning, enzyme optimization, strain engineering).
- Keep humans in the loop for execution, safety checks, and interpreting unexpected results.
- Pair AI generated ideas with automation (robots, liquid handlers) to close the loop and scale the gains.
It also reinforces the need for biosecurity and governance. AI enhanced tools can speed up benign research, but they can also lower the knowledge barrier for harmful applications. The GPT‑5 experiment was run in a benign system with tight controls, and the work was fed into a preparedness framework for managing those risks.
What’s Next for AI-Driven Biology
The 79× cloning result is a snapshot: it shows AI can reason about wet‑lab protocols, propose genuinely novel mechanisms (RecA + gp32), and combine multiple improvements into a large end‑to‑end gain. It also shows how those ideas can be tested and partially automated in a real lab, with robots achieving performance close to humans.
Looking ahead, the big wins will come from:
- Better agentic AI: Models that plan longer‑term experiments, not just single‑step optimizations.
- Tighter human‑AI workflows: Systems where scientists guide AI in plain language, and AI handles protocol design, analysis, and robot control.
- Generalization and tuning: Adapting RAPF‑HiFi and T7‑like tricks to different vectors, inserts, and cell types, and optimizing reaction times, temperatures, and enzyme concentrations.
For now, this work is a powerful reminder: AI isn’t replacing biologists; it’s becoming a new kind of lab partner, one that reads, reasons, and iterates alongside us, helping to make biology faster, cheaper, and more creative
Concluding
This blog explores how AI boosts wet-lab cloning efficiency 79×. Specifically, GPT-5 accelerated molecular biology by iteratively optimizing a standard HiFi cloning protocol through a human-in-the-loop lab setup, in collaboration with a biosecurity startup.
The result was a 79‑fold increase in cloning efficiency: the same amount of input DNA yielded 79 times more sequence‑verified clones. This gain came from two key innovations: a new enzymatic assembly method (RecA‑Assisted Pair‑and‑Finish HiFi, or RAPF‑HiFi) that uses RecA and gp32 to improve DNA pairing and annealing, and a novel transformation protocol (Transformation 7, or T7) that concentrates cells at 4 °C to boost efficiency by over 30 times.
The experiment showed that AI can propose genuinely novel, synergistic protocol changes not just minor tweaks, but fresh combinations of proteins and reaction dynamics that are not standard in current cloning workflows. To test scalability, the protocol was run on an autonomous lab robot, which matched the relative improvement seen by humans, though with lower absolute colony counts, highlighting current challenges in robotic precision.
This work fits into a broader trend: AI is moving from reading papers to designing and optimizing real lab experiments. Paired with automation and robotics, it can help close the Design Build Test Learn cycle, making biology faster, cheaper, and more engineering like. However, these advances also raise important biosecurity concerns, reinforcing the need for careful evaluation and robust safeguards in AI driven biology.
This is just one step in the journey of AI transforming science. If you’re interested in how technology is reshaping research, security, and innovation, do visit our other blog posts on our page—tagged with AI, TECH, and CYBERSECURITY—for more insights into the future of intelligent systems.





